The nucleosome core particle (NCP) constitutes the basic structural unit of the eukaryotic chromatin. It consists of 147 base-pairs (bp) of DNA wrapped in a left-handed superhelix 1.65 times around a histone octamer. The octamer is formed by two copies of the histone proteins H2A, H2B, H3, and H4. Each nucleosome is connected to it neighbors by a linker DNA (~10–80 bp). Nucleosomes can associate to form compact higher- order fiber structures (diameter of ~30 nm) that are essential for the compaction of eukaryotic DNA. The NCP structure is not static but can change in particular upon modification of histone proteins and of DNA. Regulation of gene expression, replication, and DNA repair processes are tightly coupled to transient structural changes in the chromatin and nucleosomes structures. The post-translational covalent modification includes acetylation and methylation of lysine residues and methylation of DNA. These modifications mainly occurrs on the histone tail. The modified histone tails recruit gene regulatory proteins and other signal transduction proteins either to mediate chromatin remodeling or DNA-binding properties of the histone proteins. The acetylation of lysine residues reduces the positive charge of histone proteins and may reduce the interaction with DNA. Access of buried regions in the nucleosome can be facilitated by recruitment of ATP-driven remodeling factors in response to a certain histone modification state. However, spontaneous unwrapping is also possible to transiently increase the accessibility of proteins. Fluorescence resonance energy transfer (FRET) measurements indicate that under physiological conditions, partial unwrapping of the DNA from the histone core can occur spontaneously starting from the ends of the wrapped DNA. The recently determined high resolution X-ray NCP structures are allowing a detailed understanding of the DNA and histone structure in the nucleosome and the interactions that contribute to DNA–histone complex formation. The nucleosomal DNA differs from known free B-DNA oligonucleotide structures in that it adopts a curved structure with the inner site contacting the histone octamer at several sites and the other side exposed to solvent. The nucleosome crystal structures also provided insight into the sequence dependence of the helical DNA structure and global deformability. For example, sequence-dependent differences in the mode of nucleosomal DNA-bending have been observed in the X-ray structure varying between a smooth bending and local kinking of the DNA. The distribution of distinctive sequence motifs that facilitate sharp bending and kinking of DNA is of critical importance for nucleosome formation and positioning on chromosomal DNA. Analysis of dinucleotide probability distributions in known nucleosome-binding sites has recently resulted in a genomic code for nucleosome positioning that can explain ~50% of in vivo-observed nucleosome positions. Recent experimental and computational studies on short DNA loops indicate that kinking can occur in DNA spontaneously and reversibly to allow for sharp DNA-bending. It is also not clear whether the packing strain on DNA promotes dynamic transitions of the nucleic backbone toward more noncanonical states that change the recognition properties of the nucleosomal DNA.
In collaboration with Prof. M. Zacharias (TU Muenchen, Germany), the NCP crystal structure was simulated for more than 21 ns in aqueous solution with the aim to understand the dynamic behavior of the DNA, histone proteins, and the surrounding water molecules and to get a glimpse on the mechanism of formation and stabilization [1]. The challenging simulation (~230000 atoms) was performed on the parallel cluster at the PNNL supercomputer facilities. Histone and DNA atoms stayed overall close to the crystallographic start structure. Calculated atomic and helical fluctuations showed good correlation to crystallographic B-factors. This includes a characteristic periodic flexibility pattern along the nucleosomal DNA with a much more flexible solvent exposed the side of the DNA than the side in contact with histone proteins. The core of the histone proteins showed lower fluctuations than the DNA except for the N-terminal histone tails. The histone tails underwent significant conformational rearrangements during the first 5 ns of the simulation (see Figure 1). In the X-ray starting structure, the histone tails are largely solvent-exposed contacting adjacent nucleosome particles in the crystal. During the simulations, several histone tails wrapped around the DNA modulating its accessibility The simulations allowed to characterize a number of global flexible degrees of freedom of the nucleosome core particle that may correspond to directions involved in large-scale nucleosome rearrangements during DNA recognition, transcription and replication.
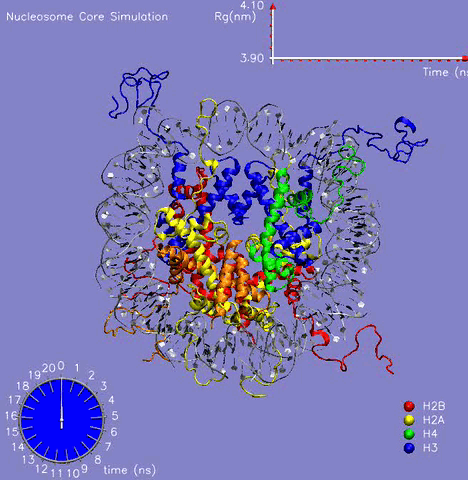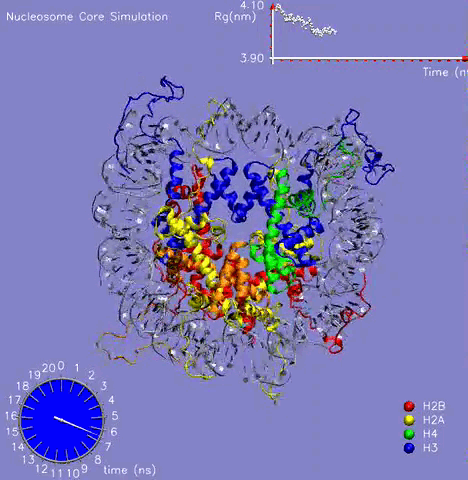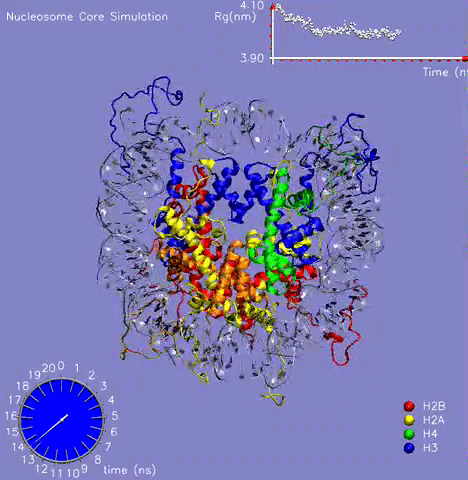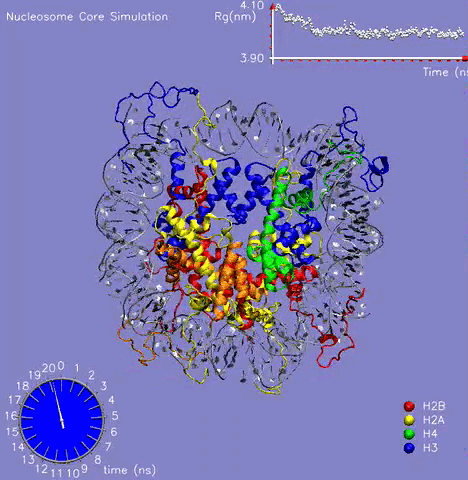
Figure 1: The 21 ns simulation of the nucleosome core particle in water. The graph on the top shows the value of the Radius of Gyration of the complex during the simulation.
MAKING OF THE FIGURE
The video animation was created using the program VMD using the following tcl script.
# the procedure disp_time draw the dials
# Draw displays for simulation time
# It was obtain from the script library
# of VMD:
# http://www.ks.uiuc.edu/Research/vmd/script_library/scripts/dials/
# Works best in orthographic projection mode!
# Author: Willy Wriggers (wriggers@ks.uiuc.edu)
proc disp_time {time} {
# radius of display in Angstrom
set radd 15.0
# radius of tube in Angstrom
set radt 0.5
# resolution of tube, number of polygon edges
set poly 100
# offset of display from center in Angstrom
set xsep 5.0
set ysep 10.0
# initialize.
set twopi [expr 4*asin(1.0)]
set pi [expr 2*asin(1.0)]
set delta [expr $twopi / $poly]
# draw rim and face color
draw color silver
set center "$xsep $ysep 0.0"
set angle1 [expr $twopi * 10.0 / 21.0]
set sx $xsep
set sy [expr $radd + $ysep]
set sz 0.0
set old "$sx $sy $sz"
set angle $delta
set i 0
while {$i <= $poly} {
set sx [expr $radd * sin($angle) + $xsep]
set sy [expr $radd * cos($angle) + $ysep]
set new "$sx $sy $sz"
draw color silver
draw cylinder $old $new radius $radt resolution 10
draw color blue
draw materials off
draw triangle $center $old $new
draw materials on
set old $new
set angle [expr $angle + $delta]
incr i
}
set sx $xsep
set sy [expr $radd + $ysep]
set old "$sx $sy $sz"
set angle $delta
set i 0
# draw hand
draw color white
set fsep [expr 2.0 * $radt]
set angle [expr ($time / 21.0) * $twopi]
set sx [expr 1.2 * $radd * sin($angle) + $xsep]
set sy [expr 1.2 * $radd * cos($angle) + $ysep]
set tx [expr -0.2 * $radd * sin($angle) + $xsep]
set ty [expr -0.2 * $radd * cos($angle) + $ysep]
draw cone "$tx $ty $fsep" "$sx $sy $fsep" radius $radt resolution 10
set radk [expr 1.5 * $radt]
draw sphere "$xsep $ysep $fsep" radius $radk resolution 10
# draw face
set angle 0.0
set angcount 0
while {$angle < $twopi - $twopi * 1.0 / 21.0} {
set sx [expr 1.1 * $radd * sin($angle) + $xsep]
set sy [expr 1.1 * $radd * cos($angle) + $ysep]
set tx [expr 0.7 * $radd * sin($angle) + $xsep]
set ty [expr 0.7 * $radd * cos($angle) + $ysep]
set radf [expr 0.6 * $radt]
draw color silver
draw cylinder "$tx $ty $sz" "$sx $sy $sz" radius $radf resolution 10 filled yes
set ux [expr 1.3 * $radd * sin($angle) + $xsep - 1.5]
set uy [expr 1.3 * $radd * cos($angle) + $ysep]
draw color white
draw text "$ux $uy $sz" "$angcount"
set angle [expr $angle + $twopi * 1.0 / 21.0]
set angcount [expr $angcount + 1]
}
#
# This is used to draw the caption
#
set sx [expr $xsep + 15]
set sy [expr -1.2 * $radd + $ysep]
draw color white
draw text "$sx $sy $sz" "time (ns)"
#
# This part draw the Legend
#
draw color blue
draw sphere " 130 0.0 $sz" radius 2 resolution 10
draw color white
draw text "135 0.0 $sz" "H3"
draw color green
draw sphere " 130 5.0 $sz" radius 2 resolution 10
draw color white
draw text "135 5.0 $sz" "H4"
draw color red
draw sphere " 130 10.0 $sz" radius 2 resolution 10
draw color white
draw text "135 10.0 $sz" "H2A"
draw color yellow
draw sphere " 130 15.0 $sz" radius 2 resolution 10
draw color white
draw text "135 15.0 $sz" "H2B"
#
# This part draw the Title
#
draw color white
draw text "-15 155.0 $sz" "Nucleosome Core Simulation"
draw color white
draw text "135 135.0 0.0" "Time (ns)"
draw color white
draw text "60 155.0 0.0" "Rg(nm)"
draw text "65 159.0 0.0" " 4.10"
draw text "65 140.0 0.0" " 3.90"
set start {80 140 0}
set end {150 140 0}
set extend [vecadd $start [vecscale 1.1 [vecsub $end $start]]]
set middle [vecadd $start [vecscale 0.9 [vecsub $extend $start]]]
graphics 0 cylinder $start $middle radius 0.50 resolution 20
graphics 0 color 1
graphics 0 cone $middle $extend radius 1.10 resolution 20
set divx [expr (70.0 / 21) ]
graphics 0 color 1
for {set i 0} {$i < 21} {incr i} {
set in [expr ($i * $divx) + 81 ]
graphics 0 cylinder "$in 139 0.0" "$in 140 0.0" radius 0.40 resolution 20
}
#
# This part draw the vertical axes of the plot
#
graphics 0 color 8
set start {81 138 0}
set end {81 158 0}
set extend [vecadd $start [vecscale 1.1 [vecsub $end $start]]]
set middle [vecadd $start [vecscale 0.9 [vecsub $extend $start]]]
graphics 0 cylinder $start $middle radius 0.50 resolution 20
graphics 0 color 1
graphics 0 cone $middle $extend radius 1.10 resolution 20
set divy [expr (20.0 / 5) ]
graphics 0 color 1
for {set i 0} {$i = 0} {
if { ! [ regexp {^[#@]} $line ] } {
set gyr($lineNumber) [lindex $line 1]
puts "$lineNumber: $gyr($lineNumber) "
incr lineNumber
}
}
close $chan
# loop through trajectory and animate
#
# production run
set offset 0.0
set scale 0.05
set num [molinfo top get numframes]
set divx [expr (70.0 / $num) ]
set divy [expr (20.0 / 0.15) ]
set sz 0.0
#set num 3
# loop through the frames
for {set i 0} {$i < $num} {incr i} {
display update off
draw delete all
# go to the given frame
animate goto $i
set time [expr $scale * $i + $offset]
set sel1 [atomselect top "segname PIH and name PG"]
set sel2 [atomselect top "segname D6"]
set coord1 [lindex [$sel1 get {x y z}] 0]
set coord2 [lindex [$sel2 get {x y z}] 0]
set restdist [veclength [vecsub $coord2 $coord1]]
disp_time $time
draw color white
for {set j 0} {$j @ stdout }]
REFERENCE
- Roccatano, A. Barthel, M. Zacharias. Structural flexibility of the nucleosome core particle at atomic resolution studied by molecular dynamics simulations. Biopolymers, 85, 401-421, (2007).
