Lignin Peroxidase (LiP), a hemoprotein isolated from the ligninolytic cultures of the white-rot fungus Phenerochaete Chrysosporium is one of the most important lignin-degrading enzymes. We have investigated the dynamical and structural properties of lignin peroxidase and its Trp171Ala mutant in aqueous solution using MD simulations [1].
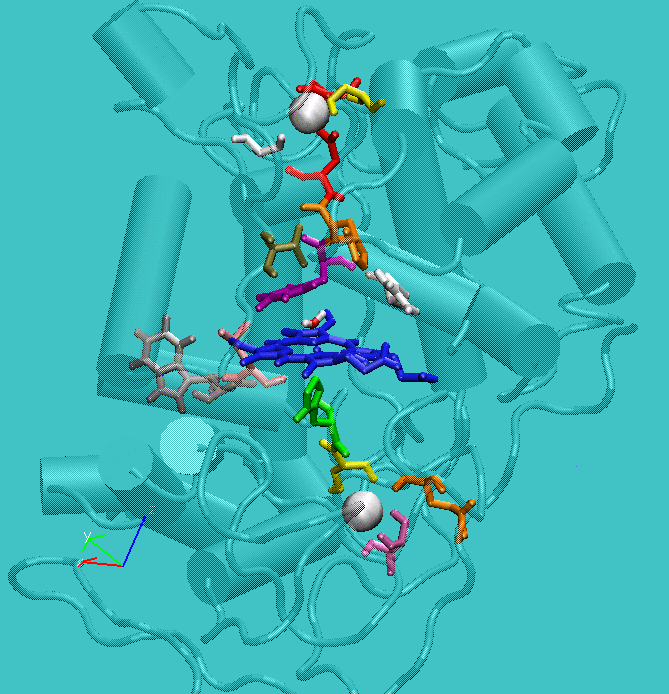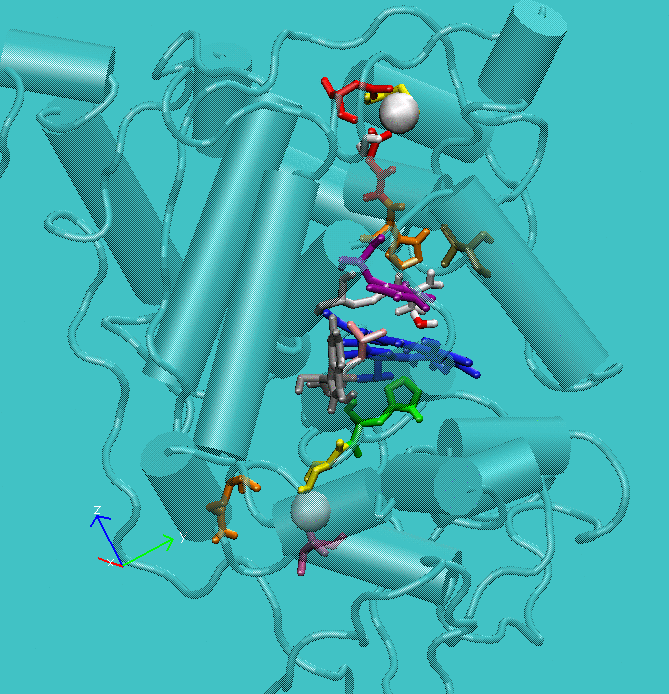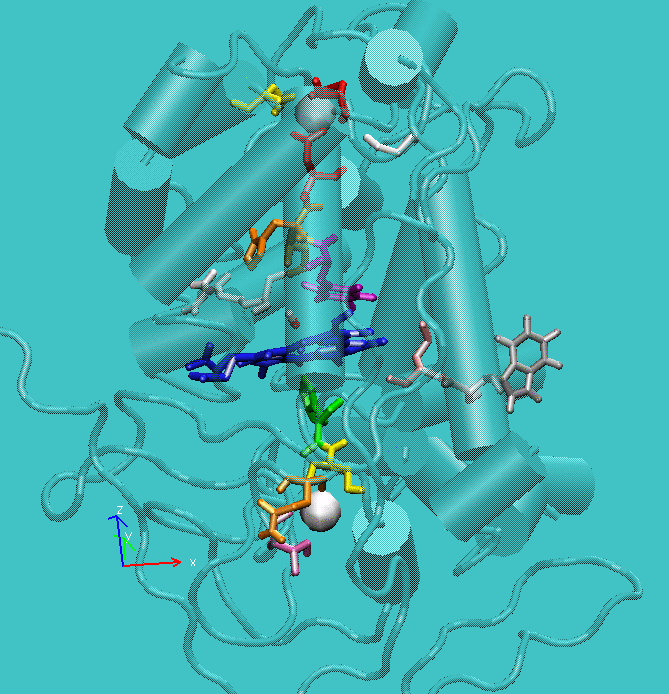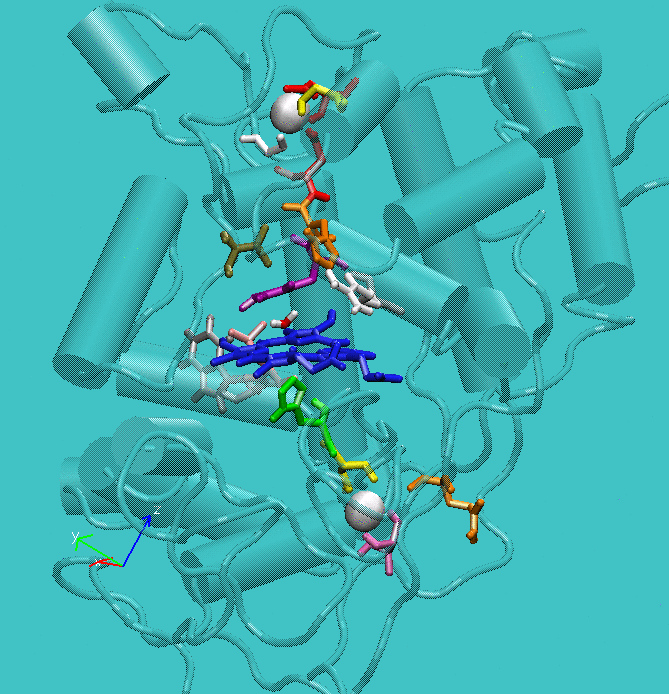
Figure 1: Structure of active site of the protein with the structural calcium ions in evidence.
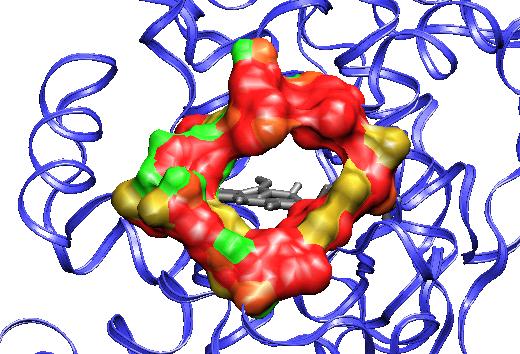
It is able to oxidize non-phenolic electron-rich aromatic compounds via an electron transfer (ET) mechanism. On the basis of the small size of both its active site and of the substrate access channel to the Heme, it is generally thought that the LiP catalyzed oxidation begins with a long-range ET followed by non-enzymatic reactions of the intermediate radical cation. In contrast, we found some pieces of evidence of a direct interaction between LiP and the substrate radical cation in the LiP catalyzed the oxidation of N, N-dimethylanilines, and aromatic sulfides. The simulations of the wild-type and the mutant enzyme have shown that in both cases the enzyme retained its overall backbone structure and all its noncovalent interactions in the course of the MD simulations. Very interestingly, the analysis of the MD trajectories showed the presence of large fluctuations in correspondence of the residues forming the heme access channel.
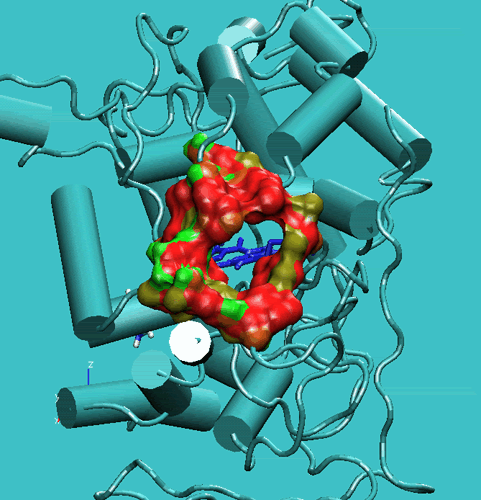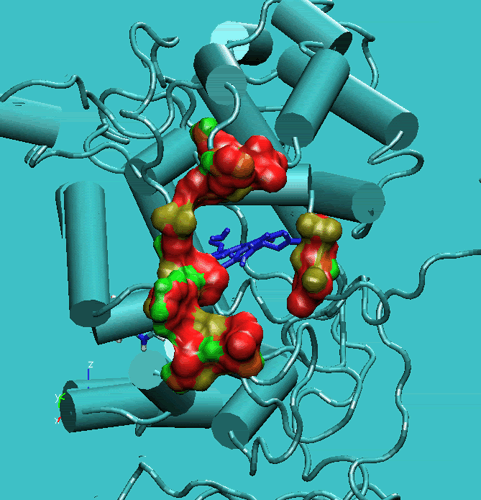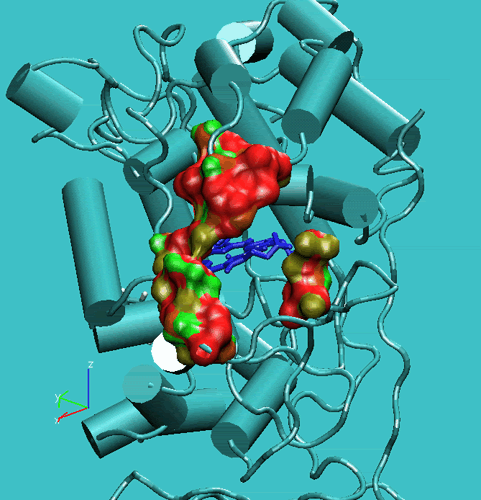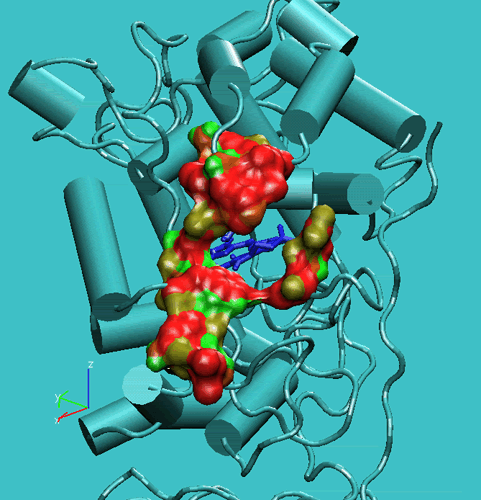
Figure 2. Access channel of the Lignin peroxidase (PDB code:1LIP). Top: in the crystal structure. Bottom: during the simulation.
These movements enlarge the opening and facilitate the access of substrates to the enzyme active site. In order to get a further insight into this respect, we have performed steered molecular dynamics docking simulations of veratryl alcohol through the access channel. The purpose of this docking study was to gain insights into the accessibility of the active site cavity by the substrate. In this respect, the substrate should be able to self-diffuse along the substrate access channel to reach the Heme edge in the active site. To increase the probability of the occurrence of such a process during the MD simulations, we have applied a harmonic force to the oxygen atom of the methoxy group of the VA molecule to pull it toward the Heme-bonded water oxygen atom (see Figure below).

Figure 3. The arrangement of for the Steered Molecular Dynamics docking of the veratric alcohol into the access channel.
In this manner, the substrate was slowly pulled through the channel up to reach a close contact (less of 0.5 nm from the VA methoxy oxygen atom) to water oxygen atom. The results of these docking simulations have shown that lignin peroxidase natural substrate (veratryl alcohol) can easily approach the Heme edge through the access channel.
REFERENCE
- F. Gerini, D. Roccatano, E. Baciocchi, A. Di Nola. Molecular dynamics simulation of lignin peroxidase. Bioph. J., 84, 3883-3893 (2003).
