In the first part of this introduction to C language, we have learnt the basic of the C language by writing simple programs for the calculation of the non-bonded interaction between two particles at variable distances. Some solutions to the first part exercises are reported in the appendix of this article.
In this second tutorial, we will learn how to use arrays data types and how to load them with a set of data read from a file. We will also use these data to perform numerical calculations and write results in output files.
Arrays and Pointers Datatypes
The program that calculate the energy of interaction between two particle doe not take in account the actual position in space of the two particle but only their distance. If we want to study the dynamics of a system composed by multiple atoms in a tridimensional space, it is way more convenient to represent the and calculate their interactions by using the coordinates directly to evaluate the distances.
Define a separate variable to describe the coordinates of many particles would be very inefficient and inconvenient. However, the C language offers a better way to deal with data sets that consist of using arrays. Arrays are the equivalent of a vector of n elements. In C language arrays are defined as:
type name[dim];
where type can be: char, int, float, double. The number of elements defines the dimension of the array and it is indicated with a value (dim) within the bracket parenthesis. The elements of the array occupy adjacent locations in memory. In C languages, arrays start at position [0].
The arrays can be accessed by referencing elements with an index or with special variables called pointers.
A pointer is a special data type variable used to store the physical memory address of a pointed variable. The syntax is:
<base type> * <pointer name>;
The memory address of a given <variable> can also be obtained using the “address operator” “&” in front of the variable name & <variable>. The content of the memory address can be obtained from the pointer variable using the (unary) dereference operator “*” of the pointer: * <pointer_variable>.
Let see in a graphical example how it works. Shall we define the variable
float vol;
and the pointer
float *pv=&vol;
with the last declaration, the memory address of the variable vol is assigned to the pointer. If the assignment is not done then it is always good practice to assign the special value of NULL (float *pv=NULL;) to avoid dandling pointers.
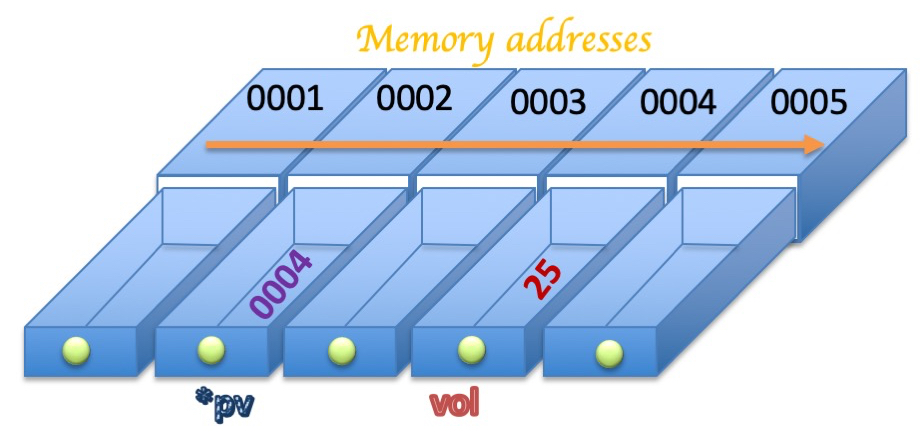
Figure 1 shows in a pictorial form the locations in computer memory (drawers) of the variable vol and its pointer *pv. Each memory location has a numerical address indicated by the number on the top.
Now, if we increase the value of vol by 5
vol=vol+5;
in the memory location assigned to this variable will change to the value of 30 (25+5). You can also increse the value of vol by using the pointer either as
vol=*pv+5;
or
*pv=*pv+5;
and now the value of vol is equal to 35.
The value of a pointer, e.g. the address, can be assigned to another pointer:
float *pq = pv;
pq now points to the same position as pv, hence, to vol, therefore
tvol = *pv + *pq
is equivalent to vol+vol.
Pointers turn to be very useful when you need to work with arrays. To understand the reason, we need to clarify the way in which the C language organize an array variable in the physical memory of the computer. Shall we define the following array for coordinates
float x[6];
the compiler will reserve 4*6 bytes in a special area of memory reserved for the storage of the program data. The memory of the computer is a grid of memory cells that the computer map and identify with a progressive number. So each physical memory location has a specific address that can be accessed by the processor to read or write variables map. The pointer datatype allow retrieve the specific memory address associated with a variable.
The pointer of the variable can be obtained by adding a star in front of the variable name. The C compiler treats the name of the array as if it were a pointer to the first element. This is important in understanding how to do arithmetic with arrays. Thus, *x is the same thing as x[0] while *(x+1) is the same thing as x[1], and so on:
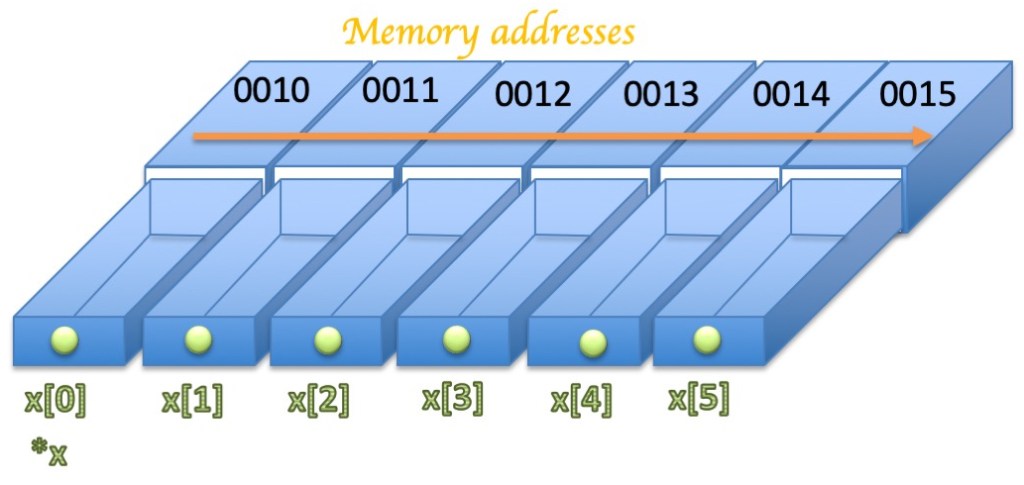
Pointers are particularly useful to pass arrays to functions.
It is now the time for you to get familiar with the use of arrays by making a simple exercise. In the following exercise, you will learn to assign the Lennard-Jones parameters (sigma and epsilon) of atomic species in Table I, to two arrays and use them to calculate the LJ potential and forces.
EXERCISE 1: Insert the following C code in the program written for the tutorial 1 (or use the one in the Appendix)
/*
The next two lineas define two arrays of three elements for
the sigma and epsilon values of LJ parameters and assign to them
initial values.
*/
float sig[3]={3.336,3.575,3.924};
float eps[3]={141.2,191.4,257.4};
float lj12[3];
float lj6[3];
float kb=6.811e-3; /* Boltzman constant X Avogadro number */
float sg,sg6,sg12;
int k;
/*
Convert the sigma and epsilon values in Table I in C12 and C6.
Units: nanometers and kJ/mol. The conversion is done on all
the three value using the k-loop. Note that the value of k
start from zero.
*/
for (k=0;k<3;k++) {
sg = sig[k]*0.1.;
sg6=pow(sg*sg,3.0);
sg12=sg6*sg6;
lj6[k]=4*eps[k]*kb*sg6;
lj12[k] =4*eps[k]*kb*sg12;
}
EXERCISE 2: Print the value of C12 and C6 on the screen.
EXERCISE 3: Define two integers (ia1, ia2) to identify the LJ set of parameters to use for the two different atoms.
Input/Output Operations
In this section, you will learn how to read and write files containing atomic coordinates, parameters or other type of data that you need for your programs. To access a file you need to open it and assign a file pointer to a special variable as in the following example:
FILE *fd;
if (!(fp=fopen(“a.xyz,”r”))){
printf (“Error opening file %s\n”, a.xyz);
}
else {
printf (“File %s is open\n”, a.dat);
}
The FILE *fd defines the file handler (fd) as a pointer (* in front of the name). All the operation that we are going to perform a file will be univocally defined by assigning the file to *fd by using the function fopen(). This also means that you can define several file handler and open more than one file at the time.
In the previous instructions a new statement is present:
If (condition) {
/* if condition is true this block of instruction is executed */
} else {
/* this block of instruction is executed if condition is false */
}
This condition statement can be to control the flow of a program according to the result of a logical condition. The logical condition is verified using logic operators (see the appendix of the first tutorial). In this case, the condition (exclamation point) is a NOT logical operation. Fopen() is a function that make accessible the file for the operation indicated in the third argument of the function. In this case the file (named: a.xyz) is open in read-only way (“r”). If you want to write data on the file then you need to specify it with the operator “w” instead of “r”. In case the file does not exist it will be created. As we will see later, fopen() returns a value that can be zero (NULL) or one. The NULL value is return by the function in case there is an error (e.g. the file does not exist or it is located in another directory). The NOT operation (!) transforms the value NULL(=FALSE) in TRUE(=1) value and direct the flow of the program in the first block of instructions, printing the message that you see in the argument of the prinf() function. If the value returned by fopen() is one than the NOT(TRUE)=FALSE and the program execute the else statement. Therefore, if-then statement allows you to make decisions in your code. We will see other example of if-else statement in other exercises.
After opening the file, you would like to read or write it. Shall we start to learn how to read a file containing for example coordinates in xyz format. The XYZ format is shown in the following example:
3 three lennard-jones particle A 0.0 0.0 0.0 A 3.1 0.0 0.0 A 0.0 3.1 0.0
The number in the first line gives the total number of atoms. The second line is a comment line. The last three lines contain in order: the atom name (for the moment let use single character names) and the x, y, z coordinates. In this case, one particle is in the origin of the reference system, the second one is along the x-axis and the third one along the y-axis. The coordinates in the XYZ format are normally given in Angstrom and you should transform them in SI units.
Copy the XYZ coordinates in a text editor and save it with the name a.xyz
To read this file we need to insert the following instructions in our program:
char ty[10];
char line[160],title[160];
float xx[10],yy[10],zz[10];
int k=0,na;
/* put here the fopen stament */
fgets (line, 80, fd);
sscanf (line,"%d",&na);
printf ("%s\n",na);
fgets (line, 80, fd);
printf ("%-80s\n",&title);
while (fgets (line, 80, fd) != NULL) {
if( line[0] != '\n' && strlen(line) > 10) {
sscanf (line,"%s%f%f%f",&ty[na],&xx[na],&yy[na],&zz[na]);
printf ("%s %f %f %f\n",&ty[na],xx[na],yy[na],zz[na]);
k++;
}
We have defined new arrays to store the information from the file. The char arrays are typically used to store alphanumeric information in C. Each element of the array contains an alphanumeric character. In this case, we have defined two char array of 160 characters, and a smaller one of 10. The first two are used to read the record (lines) of the file and to store the comment line. The numerical arrays xx, yy, zz will be used to store atomic coordinate components. After opening the file (see the previous section), the program starts to read one by one of its lines with the function fgets (line, maxb, stream). This function read the contents of the current line (record) of the stream *fd (file) for a maximum number of characters equal to maxb-1. Each character is used to fill up the array line. The function sscanf (array, format, variables)extract from the array line the contents by reading the array and assigning the variables according to the format specifies. The statement sscanf() allow extracting the atom number from the first line by reading an integer (%d) from the line. The fgets() statement assigns contents of the line to the char array title. The while-loop is then used to read the rest of the file. The condition of the while-loop is the value returned by the function fgets (). In fact, this value will be one or NULL if the record is read correctly or not. At the need of the file, fgets() return NULL and the program exit the while-loop. The if statement inside the while-loop is used to skip empty lines or lines with blanks characters.
if( line[0] != '\n' && strlen(line) > 10) {
}
The line[0] != ‘\n” checks if the first character is a newline character (empty line) and the strlen(line) checks is the length of the line is less than 10 characters (line fill with few spaces), if both the condition are true then the program read the line otherwise the line will be skipped.
Write data on a file
We use the same fopen() function to open a file for writing data but now with access operator “w” instead of “r”.
To write data on this file, you need to use the function fprinf(). This function is similar to the printf() with the difference that now you need to specify the file handler (fout) to direct the output in the output file:
Close a file
A file is closed using the function:
close(file_handler);
for example:
close(*fd);
Ones the file is closed then you can use the same file handler to open a different file.
EXERCISE 4: modify the program of EXERCISE 3 to perform the following operations:
- Read the LJ parameters from a file and assign them to arrays.
- Open a file for output writing called a.out.
- Prompt input for two numbers indicating the atom type for the calculation of the LJ interactions.
- Calculate the Lennard-Jones parameter for the interaction between the two atoms using the Berthelot-Lorentz rules. Print the sigma and epsilon parameters in the file a.out.
- Make a loop that prompt input for a distance and then calculate the LJ potential and force for the pair of atoms and the given distance. Print out the calculated values on the screen and save them also on the output file.
APPENDIX
AN EXAMPLE OF SOLUTION OF EXERCISES IN PART 1
This program calculates the energy interaction of two particles with different LJ parameters and partial charges as a function of the distance starting from a given distance and for n distance points.
#include<stdio.h>
#include<math.h>
int main() {
double V1, F1, V2, F2;
double e, s, s1, e1, s2, e2;
double s6, C6, C12, ix6, q1, q2;
double x, inc, r;
double CFAC=-9*pow(10,9);
double CFAC2=pow((1.6*pow(10,-19)),2)* 6.023*pow(10,23);
int i, n;
do{
printf("Please Enter the initial distance [nm] : ");
scanf("%lf",&x);
printf("Please Enter the distance increment [nm]: ");
scanf("%lf",&inc);
printf("Please Enter the number of sample points: ");
scanf("%d",&n);
printf("Please Enter Sigma1 (nm) : ");
scanf("%lf",&s1);
printf("Please Enter Epsilon1 (e/k) in Kelvin : ");
scanf("%lf",&e1);
printf("Please Enter Sigma2 (nm) :");
scanf("%lf",&s2);
printf("Please Enter Epsilon2 (e/k) in Kelvin : ");
scanf("%lf",&e2);
printf("Please Enter Charge on atom 1 (u.e.) : ");
scanf("%lf",&q1);
printf("Please Enter Charge on atom 2 (u.e.) : ");
scanf("%lf",&q2);
/*
* Convert in Kj/mol
*/
e1=e1*0.008312;
e2=e2*0.008312;
/*
* Apply the combination rule
*/
s=(s1+s2)/2.;
e=sqrt(e1*e2);
s6=pow(s,6.);
C6=4*e*s6;
C12=4*e*s6*s6;
r=sqrt(2)*s;
printf("\n\nSummary of LJ parameters: \n");
printf("Sigma of atom 1 = %f\n", s1);
printf("Epsilon of atom 1 = %f\n", e1);
printf("Sigma of atom 2 = %f\n", s2);
printf("Epsilon of atom 2 = %f\n", e2);
printf("Combined Sigma = %f\n", s2);
printf("Combined Epsilon = %f\n", e2);
printf("Coulomb parameters: \n");
printf("Charge on atom 1 = %lf\n", q1);
printf("Charge on atom 2 = %lf\n", q2);
/*
* Write the header of the output table
*/
printf("---------------------------------------------------------------------------------------------------------|\n");
printf(" Distance | LJ Potential Force | Coloumb Potential Force | Total Potential/Forces |\n");
printf("------------+-----------------------------+-------------------------------+------------------------------|\n");
printf(" r/nm | V(KJ/mol) F(KJ/mol/nm) | V(KJ/mol) F(KJ/mol/nm) | V(KJ/mol) F(KJ/mol/nm) |\n");
printf("------------+-----------------------------+-------------------------------+------------------------------|\n");
for(i=0;i<n;i++){
ix6=1/pow(x,6);
V1=C12*ix6*ix6 - C6*ix6;
F1=(12*C12*ix6*ix6 - 6*C6*ix6)/x;
V2= CFAC*(q1*q2/x)*CFAC2;
F2= CFAC*(q1*q2/pow(x,2))*CFAC2;
/*
* Write the results in tabular form
*/
printf(" %lf | %e %e | %e %e | %e %e \n", x,V1,F1,V2,F2,V1+V2,F1+F2);
x+=inc;
};
printf("Type zero or a negative number to exit: ");
scanf("%lf",&x);
}while(x>0);
}
