Sphere. From Space, from Space, Sir: whence else?
Square. Pardon me, my Lord, but is not your Lordship already in Space, your Lordship and his humble servant, even at this moment?
Sphere. Pooh! what do you know of Space? Define Space.
Square. Space, my Lord, is height and breadth indefinitely prolonged.
Sphere. Exactly: you see you do not even know what Space is. You think it is of Two Dimensions only; but I have come to announce to you a Third — height, breadth, and length.
Square. Your Lordship is pleased to be merry. We also speak of length and height, or breadth and thickness, thus denoting Two Dimensions by four names.
Sphere. But I mean not only three names, but Three Dimensions.
Adapted from:
Flatland: A romance of many dimensions by Edwin A. Abbott
ADVENTURE IN SPACELAND
In part 2 of this tutorial, we have learned how to use arrays and how to read atomic coordinates from a file. In the appendix, you can find an example of the solution to the exercises given in the previous tutorial.
In this third part, we are going to learn how to generate three-dimensional coordination of atoms in a cubic crystal lattice and how to calculate non-bonded molecular potential and the force acting among them.
For this purpose, we need to modify the subroutines for the calculation of one-dimensional Lennard-Jones potentials and their forces developed in the previous tutorial to account to use directly the atomic coordinate of particles located in a tridimensional space.
The Cartesian distance of two points in a Euclidean space is given by:
where and
are the coordinates of the two point particles. This expression will be used to modify the LJ energy potential and the components of the atomic forces. The latter are derived taking in account the vectorial nature of the forces in the following components:
these expressions give the components of forces acting on the atom i. The force on the atom is given by , by Newton’s third law:
Lex. III. Actioni contrariam semper et equalem esse reactionem: sive corporum duorum actiones in se mutuo semper esse aequales et in partes contrarias dirigi.
Isaac Newton. Philosophiae Naturalis Principia Mathematica. London, 1686
Finally, the square root function in C is calculated by the sqrt().
EXERCISE 2: modify the program in part 2 of this tutorial to include the possibility to read a file containing the coordinated in xyz format of two LJ particles. Using those coordinates, calculate the potential and forces acting between the two particles. Change the value of the coordinates and report the same value for different distances.
ADDING AN ELECTRIC FIELD
Atoms in molecules are surrounded by an electron density that depends on their tendency to delocalize their electron (these properties are usually referred to as electronegativity). The electron distribution produces an electric field around the molecule that can be matched by assigning partial electric point charges to each atom (I have explained in another article how this is accomplished). Therefore, we can add to the expression of the LJ potential a Coulomb potential term that describes the electrostatic pair interactions of charged atoms.
The force components for the Coulomb term on the particle i interacting with the particle j are given by the expressions
The are the partial charges and
the dielectrioc constant of the vacuum and of the medium. The force on the atom is given by the Netwon’s third law. To extend the calculation to many-particles systems, we need to perform a double sum is used to calculate all the pair-wise interactions as follows
The index i>j means that in the second sum we have to exclude all the pair already accounted by the first index. If we have four particles for example we need to calculate the following 4*(4-1)/2=6 pairs
i: 1 1 1 2 2 3
j: 2 3 4 3 4 4
when the index i=2 the index j start from 3 since 2-1 has been calculated and 2-2 does not make sense.
To program this double summation, we need to use two nested loops with two indexes (i and j). The following pseudo-code shows how to make the double loops. The potential and forces are updated using the unary operators += and they need to be initialized to zero. The results will be printed at the end of the calculation using a loop on the number of atoms (na).
Vt=0; /* total potential */
for (i=0;i<na;i++) {
F[i]=0.0;
V[i] =0.0; } /* initialize the forces */
for (i=0;i<na;i++) {
for (j=i+1; j< na; j++) {
calculate the distance using xx[i],xx[j],yy[i],yy[j],zz[i],zz[j]
calculate the LJ potential and forces
V+= V[i] + ….
Vt+= V[i] + ….
F[i]+= F[i] + …
}
}
/* Add here the printout of the potential for each atom and the total potential */
EXERCISE 3: implement the double loop to calculate the potential energy and forces for the coordinates generated by the program from EXERCISE 1.
CUBIC CRYSTAL LATTICE COORDINATES GENERATION
Atomic coordinates for simple regular systems, such as a cubic crystal lattice, can be easily generated using nested for-loops. Let start with the example of a simple one-dimensional lattice, where the atoms are lined up along the x-axis. This means that the coordinates of these atoms along the other two directions are null. The coordinate of a given atom along the x-direction is generated by multiplying the lattice index for the lattice size. The following piece of code shows how to implement the algorithm using for-loops. The variable x-axis is the lattice space in Angstrom units and nx is the number of lattice points along the x-direction.
xaxis = 3.0; /* this value can be given in input */
nx = 5;
xx[0] = 0;
for (i=1;i<=nx;i++) {
xx[i] = i*xaxis;
}
We can extend the method for a 2 D lattice using a nested for-loop as in this code snippet:
xaxis = 3.;
yaxis = 3.;
nx=5;
ny=5;
xx[0] = 0.;
yy[0] = 0.;
for (i=0;i<nx;i++) {
xx[i] = i*xaxis;
for (j=0;j<ny;j++) {
yy[j] = j*yaxis;
}
}
Now putting together what we have learned from the last tutorial, we can write a code that generates the coordinates of a squared two-dimensional lattice and save them in xyz format. The code is reported in the appendix. The coordination in xyz format (filename out.xyz) can be visualized them using the program RasMol. If you run from the command line type RasMol –xyz 2dlattice.xyz.
With nx=10, and ny=1 you should get the 1D-dimensional lattice that is visualized with RasMol (background white, space fill) will look like the following picture
With nx=10, and ny=10 you should get the 2D-dimensional lattice that is visualized with RasMol (background white, ball&stick) will look like the following picture
EXERCISE 1. Extend the 2D lattice program to calculate coordinates of atoms in a 3D lattice for a given number of lattice points along the three axes (nx, ny, nz) and for a given lattice sizes (xaxis, yaxis, zaxis). You may want also to add the possibility to input these parameters from the keyboard to make the program more user-friendly.
APPENDIX
TABLE I: LENNARD-JONES PARAMETERS FOR NOBLE GASES
| Gas | ( | |
| Ar | 0.3336 | 141.2 |
| Kr | 0.3575 | 191.4 |
| Xe | 0.3924 | 257.4 |
PROGRAM I: Two dimensional cubic lattice generator
/*
PROGRAM NAME : 2D reticulum generator
DESCRIPTION : the program generate one or two dimensional
lattices having nx,ny alternating atoms in the x
and y direction, respectively. The atoms are separated
ati equal distances xsize,ysize
number of atoms in each dimension to be set
AUTHOR : Danilo Roccatano
DATE : BREMEN, 2007
*/
#include <stdio.h>
#include <math.h>
#include <string.h>
#include <unistd.h>
int main() {
char sym1[]="Na";
char sym2[]="Cl";
char csym[3];
float xaxis,yaxis;
int nx,ny;
int nn,n,nsel;
float xx[10];
float yy[10];
float zz;
/*
* Initialize variables
*/
xaxis=2.82;
yaxis=2.82;
nx=5;
ny=5;
zz=0.0;
/*Opening output file*/
FILE *fout;
if (!(fout=fopen("2dlattice.xyz","w"))){
printf ("Error opening file %s\n", "2dlattice.xyz");
}
/*Writing output header*/
n=nx*ny;
fprintf(fout,"%i\n",n);
fprintf(fout,"%i %s%s-atoms in two dimensional rectangular lattice\n",n,sym1,sym2);
nn=0;
/*Computing and writing the lattice coordinates*/
for(int i=0;i<nx;i++){
xx[i]=i*xaxis;
for(int j=0;j<ny;j++){
yy[j]=j*yaxis;
nn+=1;
if (nn%2) strcpy(csym, sym1);
else strcpy(csym,sym2);
fprintf(fout,"%2s %8.3f %8.3f %8.3f %5d\n", csym, xx[i], yy[j], zz,nn);
}
}
printf("Generated a rectangular lattice of %i %s%s-atoms. \n",n,sym1,sym2);
fclose(fout);
}
AN EXAMPLE OF SOLUTION FOR THE PART 2
This is an example of the solution to the exercises for the last practical. The program read the parameters from the file “ljparam.dat” containing the following data.
LJ parameters 3 Atom_name sigma(A) epsilon(K) Ar 3.336000 141.200000 Kr 3.575000 191.400000 Xe 3.924000 257.400000
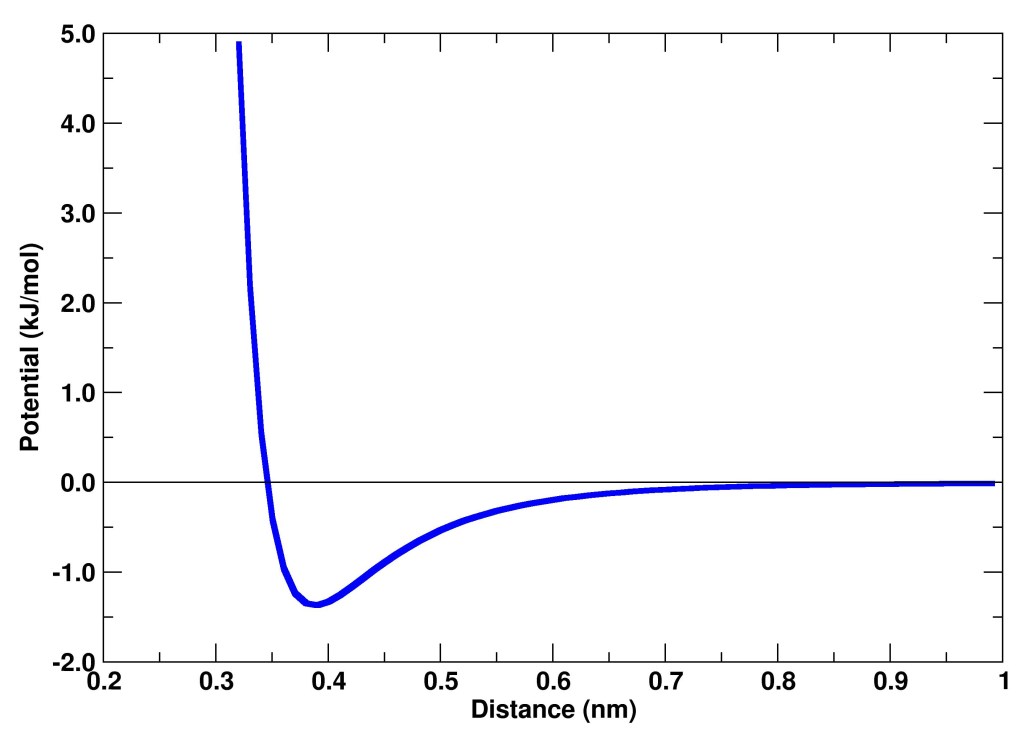
The program asks to choose two atoms types, then it calculates the combined parameters and asks for the maximum value of distance to calculate the value of the potential and the force. The values are stored in the file “Results.out“, and they can be plotted with xmagrace (see the graph at the end).
#include <stdio.h>
#include <math.h>
#include <string.h>
int main() {
char line[160],title[160];
char aty[10];
char yn[2];
int k=1;
int ia1,ia2;
int np; // number of parametes in the library
float V,s,s1,s2,e,e1,e2,F;
float x,rmin,xmin,xmax,xinc;
float C6,C12;
float s6,ix6,ix;
float sig[10];
float eps[10];
float kb=8.3145e-3; /*Boltzmann constant*Avogadro Number */
FILE *fd;
FILE *fout;
/*
* READ LJ PARAMETERS FROM A LIBRARY FILE
*/
printf("\nREADING THE LJ PARAMETER FILE \n");
if (!(fd=fopen("ljparam.dat","r"))){
printf("\nError opening file %s\n", "ljparam.dat");
}
fgets (title, 80, fd);
fgets (line, 80, fd);
sscanf (line,"%d",&np);
fgets (line, 80, fd);
printf (" %s \n",line);
while (fgets (line, 80, fd) != NULL){
if (line[0] != '\n' && strlen(line) > 10) {
sscanf (line,"%s%f%f",&aty[2*k],&sig[k],&eps[k]);
printf ("%i %s %8.3f %8.3f\n",k, &aty[2*k],sig[k],eps[k]);
k++;
}
}
fclose(fd);
/*
* OPEN THE OUTPUT FILE
*/
if (!(fout=fopen("Results.out","w"))){
printf ("Error opening file %s\n", "Results.out");
}
/*Input for the atom types*/
printf("\n");
printf("Enter the atomtype for particle 1: ");
scanf("%i",&ia1);
printf("Enter the atomtype for particle 2: ");
scanf("%i",&ia2);
printf("\n");
/*Berthelot-Lorenz mixing, etc.*/
s=0.5*(sig[ia1]+sig[ia2])/10.;
e=kb*sqrt(eps[ia1]*eps[ia2]);
s6=pow(s,6);
C6=4*e*s6;
C12=4*e*s6*s6;
rmin=pow(2.,1./6.)*s;
/*Output the results on the screen*/
printf("Mixed Lennard-Jones parameters:\n");
printf(" Epsilon: %f kJ/mol \n",e);
printf(" R min : %f nm \n",rmin);
printf(" Sigma : %f nm\n",s);
printf(" C6 : %f kJ*nm^6/mol \n",C6);
printf(" C12 : %f kJ*nm^12/mol \n\n",C12);
/* Estimate the minimum distance to calculate the LJ function */
xmin=s-0.025;
ix=1./xmin;
ix6=1./pow(xmin,6.);
V=C12*ix6*ix6-C6*ix6;
printf(" Value of Potential V(xmin=%f nm): %f kJ/mol \n",xmin,V);
/*Output the results in the file */
fprintf(fout,"#Lennard-Jones parameters:\n\n");
fprintf(fout,"# Atom Type 1 : %c%c \n",aty[2*ia1],aty[2*ia1 + 1]);
fprintf(fout,"# Sigma : %f nm/10\n",sig[ia1]);
fprintf(fout,"# Epsilon : %f K\n\n",eps[ia1]);
fprintf(fout,"# Atom Type 2 : %c%c \n",aty[2*ia2],aty[2*ia2 + 1]);
fprintf(fout,"# Sigma : %f nm/10\n",sig[ia2]);
fprintf(fout,"# Epsilon : %f K\n\n",eps[ia2]);
fprintf(fout,"# Mixed Sigma : %f nm\n",s);
fprintf(fout,"# Mixed Epsilon: %f kJ/mol \n",e);
fprintf(fout,"# C6 : %f kJ*nm^6/mol \n",C6);
fprintf(fout,"# C12 : %f kJ*nm^12/mol \n\n",C12);
fprintf(fout,"# Distance [nm] Potential [kJ/mol] Force [nN]\n");
/*Input for the value of the maximum distance range*/
do {
printf("\nEnter the value of the maximum distance distance (in nm): ");
scanf("%f",&xmax);
printf("\nEnter the distance increment (in nm): ");
scanf("%f",&xinc);
printf("Finish?(y/n)\n");
scanf("%s",yn);
} while (strcmp(yn,"y"));
for (x=xmin;x<=xmax;x+=xinc){
/*Calculate the LJ potential and the force between xmin and xmax x*/
ix=1./x;
ix6=1./pow(x,6.);
V=C12*ix6*ix6-C6*ix6;
F=(12.*ix*C12*ix6*ix6-6.*ix*C6*ix6)/602.2; /*to obtain [nN]*/
fprintf(fout," %f %f %f\n", x,V,F);
}
fclose(fout);
return 0;
}