Understanding the conformational transitions that trigger the aggregation and amyloidogenesis of otherwise soluble peptides at atomic resolution is of fundamental relevance for the design of effective therapeutic agents against amyloid-related disorders. I have collaborated to study the transition from ideal α-helix to ß-hairpin conformation by long time scale, all atoms molecular dynamics simulations in explicit water solvent [1]. Two amyloidogenic peptides have been investigated: the H1 peptide from prion protein and the Aß(12-28) fragment from the Aß(1-42) peptide responsible for Alzheimer disease.

Figure 1: Summary of the structural behaviour of the H1 peptide from Prion Protein.

Figure 2: Experimental data available (before 2004) on the peptide H1.

Figure 3: Summary of the MD simulations of the H1 peptide.
The MD simulation starting from the low-resolution X-ray structure (see Figure I) is reported in Figure 4. A stable beta-haiprin is formed within 8 ns.
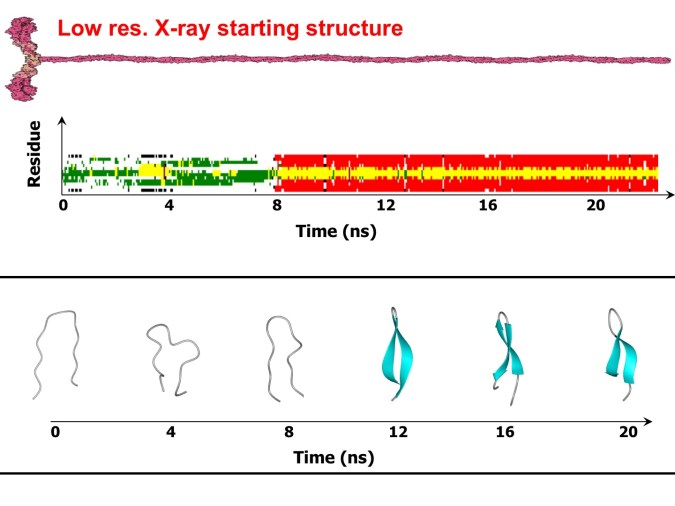
Figure 4. MD simulation at 300 K starting from the low-resolution X-ray structure. The beta-hairpin is formed at t = 8 ns.
The simulations evidence the unfolding of α-helix, followed by the formation of bent conformations and a final convergence to ordered in register ß-hairpin conformations (see Figure 4).
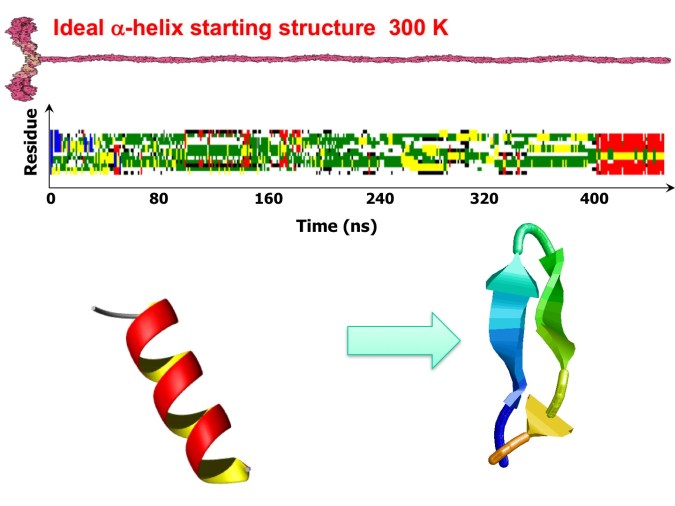
Figure 4. Top. Time evolution of the H1 peptide secondary structure. The analysis was performed with the DSSP program. MD simulation at 300 K starting from an ideal -helix. Note the formation of the beta-hairpin at t = 408 ns. Bottom. The starting and final structures of each simulation are shown on the left and right sides, respectively.
The ß-hairpins observed, despite different sequences, exhibit a common dynamic behaviour and the presence of a particular pattern of the hydrophobic side chains, in particular in the region of the turns. These observations hint at a possible common aggregation mechanism for the onset of different amyloid diseases and a common mechanism in the transition to the ß-hairpin structures.
Furthermore, the simulations presented herein evidence the stabilization of the Alpha-helical conformations induced by the presence of an organic fluorinated co-solvent.
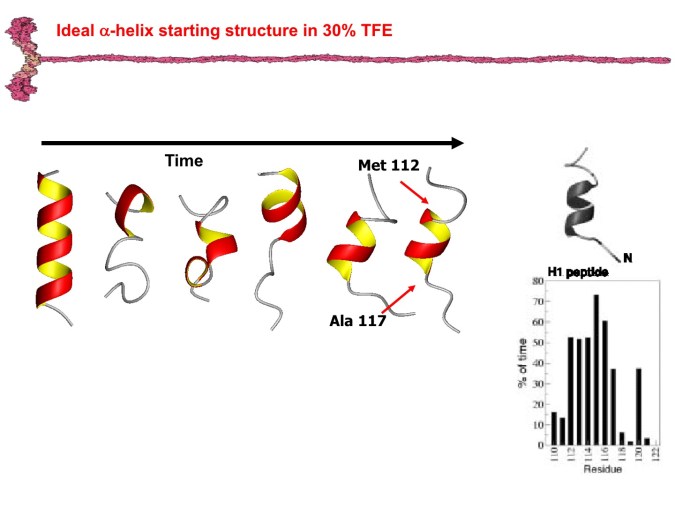
Figure I. MD simulation at 300 K starting from H1 in ideal alpha-helix in 30% TFE. The alpha-helix structure remains mainly preserved as from the experiments.
The results of molecular dynamics in 2,2,2-trifluoroethanol (TFE)/water mixture provide a further evidence that the peptide coating effect of TFE molecules is responsible for the stabilization of the soluble helical conformation.
REFERENCES
- Daidone, F. Simona, Roccatano, R. A. Broglia, G. Tiana, G. Colombo, A. Di Nola. β-hairpin conformation of fibrillogenic peptides: structure and α/β transition revealed by molecular dynamics simulations. PROTEINS: Struct., Funct. and Bioinf., 57, 198-204 (2004).
