“To see a World in a Grain of Sand
And a Heaven in a Wild Flower,
Hold Infinity in the palm of your hand
And Eternity in an hour.”
―William Blake, Auguries of Innocence.
INTRODUCTION
This blog is based on public talks that I have delivered at the University of Lincon, UK and at the Gravity Fields Festival 2016. Here I give a short summary of talk topics.
Nature is an unlimited source of great inspiration (and imitation) for scientist and engineering. In fact, the continuous advance in the knowledge of the complex machinery of life is producing profound impacts in the modern societies. Life, in the form that we know, definitively exploited what we now call “nanotechnology” to emerge. Living cells are crowded with fascinating molecular machines with a large variety of functions not yet completely explored. Nature as a blind and patient engineer builds these machines without a defined blueprint but utilizing the power of the evolution.
In the last 50 years, we have made important progress in the understanding the principle governing the mechanism of function of the nanometer-sized engineering marvels. In this primer, I will make a voyage to explore these nuts and bolts of our cell. You will discover that in our body there are machines having functions similar to those used in our day-life experience. You will also discover that it is possible to study their mechanism and function by using the same basic physical laws discovered 350 years ago by the universal genius Isaac Newton. Namely, the same principles that describe the motion of stars in our galaxy are helping us to unravel the complex machinery of life.
A fantastic voyage
The title of this blog is inspired to the famous 1966 science-fiction movie Fantastic voyage (see https://www.imdb.com/title/tt0060397), popularized with the novel written by the polymath science fiction writer Isaac Asimov. Unfortunately, we did not yet discover the technology* that in the movie allowed to miniaturize a submarine and its crew to the size of a cell. So we do not have the possibility to feel the molecules that we are going to explore for real! However, we can still use as submarine our imagination and the power of computer graphics to have fantastic visual experience.
* In the sequel novel Fantastic Voyage II: Destination brain (1987), I. Asimov gave a plausible scientific ground to the miniaturization technology used to reduce the size of the submarine. He imagined that by altering one fundamental constant governing the subatomic world, the scale of objects could be changed. By using the words of Isaac Asimov:
“Why are you so certain miniaturization is impossible?”
“If you reduce a man to the dimensions of a fly, then all the mass of a man would be crowded into the volume of a fly. You’d end up with a density of something like –” he paused to think — “a hundred and fifty thousand times that of platinum.”
“But what if the mass were reduced in proportion?”
“Then you end up with one atom in the miniaturized man for every three million in the original. The miniaturized man would not only have the size of a fly but the brainpower of a fly as well.”
“And if the atoms are reduced, too?”
“If it is miniaturized atoms you are speaking of, then Planck’s constant, which is an absolutely fundamental quantity in our Universe, forbids it. Miniaturized atoms would be too small to fit into the graininess of the Universe.”
“And if I told you that Planck’s constant was reduced as well, so that a miniaturized man would be encased in a field in which the graininess of the Universe was incredibly finer than it is under normal conditions?”
The miniaturization field devised by Asimov in his novel was based on the idea of reducing the Planck constant h. Indeed, if we look to the classical formula describing the size of a microscopic object as the radius of the orbit of electrons around the proton in the hydrogen atom in the Bohr formulation: , we can clearly see the linear dependence of a0with h. However, we do not know if this will be ever possible to realize in practical and, if it can be done, how it can be realized. Fortunately, we do not need to be so small to explore the microscopic world within us. We can use microscopes to look to this world from the less adventurous but definitively more comfortably view of our computer screen.
For those of you that have the possibility to access VR gears, I recommend giving a look to the program Haptimol (http://www.haptimol.com) that my friend Dr. Steven Hayward and his colleagues are developing at the University of East Anglia, UK. The program can give an immersive experience and if you use the program eating a steak you can get the complete sensitive virtual experience of protein molecules!
IT WILL BE CONTINUED…STAY TUNED!
In my talk at the Gravity Fields Festival 2016, I have shown different models of molecular machines, I have added some of them to this blog with the details of their construction. The models have been generated using a small script in awk language and represented using the program VMD (http://www.ks.uiuc.edu/Research/vmd/).
The Photosynthetic Apparatus of Rhodospirillum photometricum
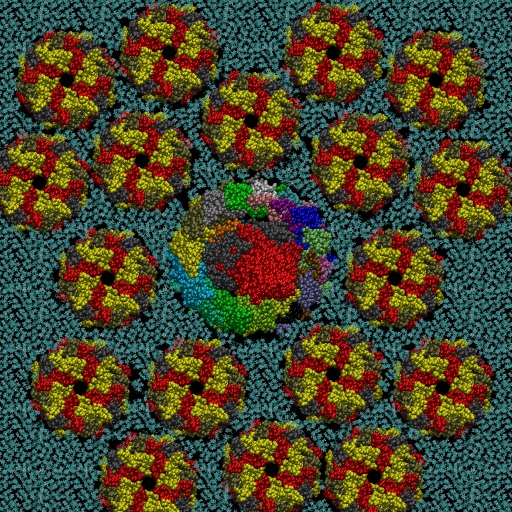
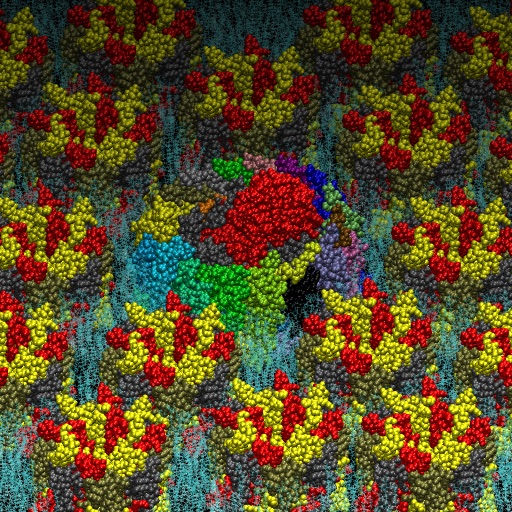
The photosynthetic apparatus of purple photosynthetic bacteria is particularly simple and it is located in a specialized membrane system that develops in the bacterial cytoplasm. They are composed by four integral membrane protein complexes: a peripheral LH2 antennae complexes that serve to collect light and transfer the absorbed energy to the second complex, the core-complex, which is constituted of an antenna complex (LH1) associated with the photochemical reaction center (RC). The LH1 serves to funnel the light energy to the RC where a charge separation takes place catalyzing the oxidation of a water-soluble carrier. Different crystallographic structures of these components are available in the Protein Data Bank. Based on the supramolecular assembly of this protein observed using Atomic Force Topographic (Scheuring et al. 2007), it was possible to generate a molecular model of the photosynthetic apparatus off Rsp. photometricum. The model comprises one RC-LH1 core complex surrounded by several LH2 complexes. The model is a chimera of crystallographic structures from different organisms. In particular, for the LH2 was used the nonameric Rps. acidophila structure [PDB id: 1KZU (McDermott et al., 1995)].
The LH1-RC complex is based on the recent crystal structure from Thermochromatium tedium [PDB id: 4V8K (Niwa et al. Nature, 2014)]. LH2 and the core complex were first centered and aligned and then translated and rotated to their relative positions using as reference the AFM topography reported by in Scheuring et al. 2007. A DPPC lipid bilayer was also generated using the tools in the VMD program and added to the model by removing the lipid molecules overlapping with the proteins.
Two views of the model are reported below. The complete fly-by animation is here.
The Yeast V-ATPase
Vacuolar-ATPase (V-ATPase) is an ancient enzyme with remarkably diverse functions in eukaryotic organisms. It is used to acidify different organelles and as a proton pump across the plasma membranes of numerous cell types. For its functions, V-ATPases use the energy produced by ATP hydrolysis. It is generally seen as the polar opposite of ATP Synthase that uses the energy from a proton gradient to produce ATP.
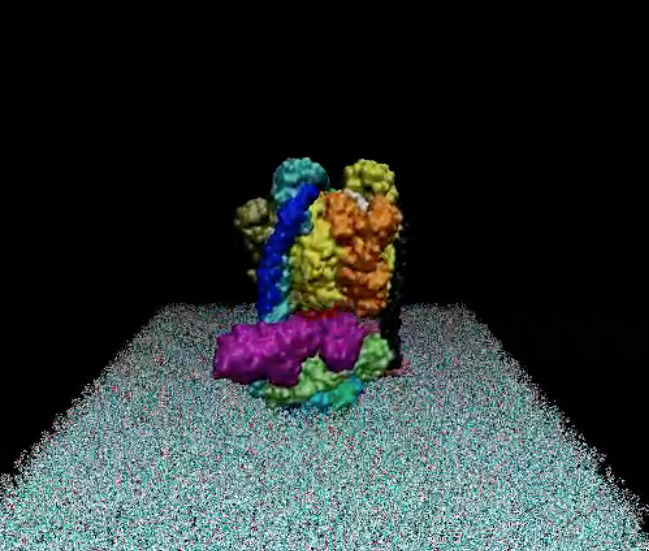
The structure of the model of the Yeast V-ATPase was generated by Zhao et al. using electron microscopy add homology modeling (PDB Id: 3J9U, Zhao et al. Nature, 2015). I have just generated a DPPC lipid bilayer using the tools in the VMD program and just added to the model by removing the lipid molecules overlapping with the proteins.
A view of the model is reported below. The complete fly-by animation is here.

Hi Paul,
thank you for reading my blog. I understand your diappointment as I didn’t have time to add more information. I am planning to update it adding the presentation slides and some reference to the topics.
LikeLike