After a long pause, the adventure in the PERL programming language series continues with another example of biophysics and molecular biology science application.This time, we are going to make a program to model the effect of the temperature on DNA stability in solution.
The effect of temperature DNA structure integrity plays an important role in molecular biology applications. For example, the DNA amplification method based on polymerase (the PCR method) is based on a series of temperature cycles to separate the two strand of DNA to replicate. DNA primers are used to initiate the process and the knowledge of their melting temperature (Tm) play an important role in optimizing the DNA amplification process.
The first attempts to create a model of DNA thermodynamics date back to the beginnings of 1960. Studies pionered by the groups of Zimm [1] and Tinoco [2], have shown that the relative stability of a double-stranded DNA molecule depends primarily on the nature of the nearest-neighbor bases along the sequence. This finding brought to the formulation of a simple mathematical model (called the nearest-neighbor (NN) model) to predict relative stabilities of double stranded DNA according to the nucleotide sequence [see Cantor]. Subseguently, the NN model was further improved by the contribution of several research groups. In particular, Santalucia and co-workers. proposed a set of parameters for the NN model that provides an excellent prediction of the thermodynamic properties of short DNA homo oligonucleotides and are commonly used to calculate the stability of DNA primers used for PCR applications.
In this third article on PERL programming, I will give some indication on how to implement a simple version of the NN model. To make more fancy the programming task, we are also going to provide the program with a simple Graphical user interface using PERL/Tk.
The NN model
NN model is based on the pairwise interactions between adjoint pair of nucleotides along a DNA sequence. Taking in account the four nucleotides (A, C, T, G) composing the DNA, ten different NN pairwise interactions are possible in a double stranded DNA molecule. These pairwise interactions are AA/TT, AT/TA, TA/AT, CA/GT, GT/CA, CT/GA, GA/CT, CG/GC, GC/CG, and GG/CC for strand directions and the bar indicate the complementary pair. In addition, there is an additional contribution determined by the starting pair contributed to the total free energy of the double strand interaction. Finally, the C2 symmetry in the DNA chain play also a role that has been quantified with additional correction term.
The thermodynamics values of enthalpy and entropy and Gibbs free energy difference for each of the nucleotide pair interactions have been derived from experimental measurements. Santamaria et al. have proposed a set of parameters by analyzing the different values obtained by several groups. This unified set of parameters is geenrally used for the the model. In the following table, the Santalucia et al. parameters are reported.
| Nearest-Neighbor nucleotides pairs | |||
| AA/TT | -1.00 | -7.9 | -22.2 |
| AT/TA | -0.88 | -7.2 | -20.4 |
| TA/AT | -0.58 | -7.2 | -21.3 |
| CA/GT | -1.45 | -8.5 | -22.7 |
| GT/CA | -1.44 | -8.4 | -22.4 |
| CT/GA | -1.28 | -7.8 | -21.0 |
| GA/CT | -1.30 | -8.2 | -22.2 |
| CG/GC | -2.17 | -10.6 | -27.2 |
| GC/CG | -2.24 | -9.8 | -24.4 |
| GG/CC | -1.84 | -8.0 | -19.9 |
| Initiation at A-C | 0.98 | 0.1 | -2.8 |
| Initiation at A-T | 1.03 | 2.3 | 4.1 |
| Symmetry correction | 0.4 | 0 | -1.4 |
To understand how to use these parameters in the NN model, I will show an example of calculation for a short homonucleotide DNA sequence.
Let consider the following double strand DNA sequence:
5′-CGTACG-3’/3′-GCATGC-5′
Using the Table 1, we sum up the contribution of the NN nucleotides as follows
We need to add the contribution from the terminal nucleotide
.
We also notice that the sequence is symmetric, therefore we need add a symmetry contribution term
Finally, the total free energy of dissociation is obtained adding all the contributions
Using the same procedure, we also calculate the enthalpy and the entropy of dissociation obtaining
The melting temperature. At the melting temperature only half of the DNA double strands in a solution are complementary base-paired. To estimate its value, we assume that the unfolding process for short DNA strands can be approximated as a two-state model. In this approximation, we have that
where is the concentration of the DNA in solution (in our case we set the value to
).
This equation gives the value of the temperature in Kelvin and for a 1 M sodium ions concentration. However, it is possible to improve the accuracy of the Tm value by taking in account the effect of ions and denaturants on the equilibrium.
The population of the folding/unfolding population can be calculated from the folding/unfolding free energy, using the relation
that generate a typical sigmoidal shaped curve.
The implementation in Perl/Tk
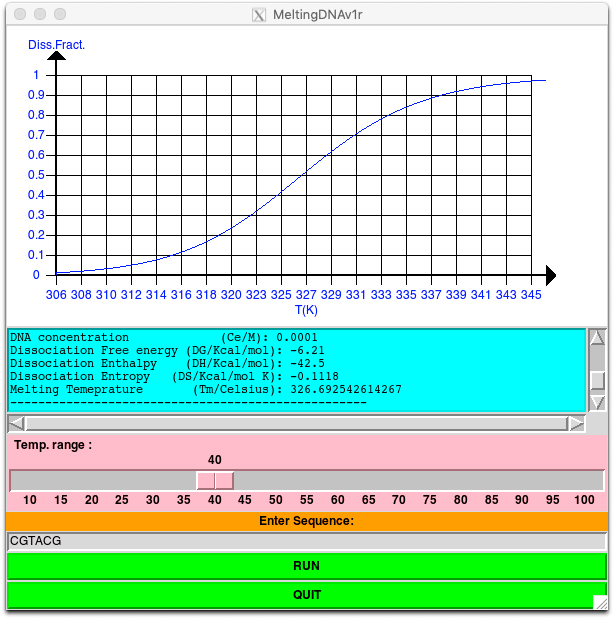
The PERL/Tk code reported in the appendix is an implementation of the model described in the previous paragraphs. In the Tk GUI, you can insert the nucleotide sequence and the range of temperatures with respect the Tm to visualize for the population curve. The concentration of the DNA is fixed to 0.0001 M. You can modify this value in the program but changing the variable Ce. Upon runnign the code, you can read the value of free energies, enthalpies and entropies and the Tm. In addition, the population curve is plotted in the upper panel as shown in the Figure 1.
Note that the reported program is provided for sole educational purposes. It is possible to modify, correct and improve the program to the condition that the resulting code is made publicy available.
References
- Crothers,D.M.&Zimm,B.H. J.Mol.Biol. 1964. 9, 1-9.
- DeVoe H, Tinoco I., Jr J Mol Biol. 1962, 4, 500–517.
- Hatim T. Allawi and John SantaLucia, J. Biochemistry. 1997, 36, 10581-10594.
APPENDIX
MeltingDNA: a Perl/Tk to calculate the melting curve of short DNA homooligonucleotides.
#! /usr/bin/perl -w
#
# PROGRAM: MeltingDNA.pl
#
# Author daniloroccatano <danilo dot roccatano at gmail.com>
# Version 1.0
# Copyright (C) 2018-2021 Danilo Roccatano
#
use strict;
use Tk;
use Tk::FileSelect;
use Tk::Scale;
my $Ce=0.0001;
my $R=0.001986;
#
# DNA THERMODYNAMMICS CALCULATIONS
# DNA Melting temperature using the Nearest-neighbor sequence's method
#
# Parameters from
# John SantaLucia Jr. (1998). "A unified view of polymer, dumbbell,
# and oligonucleotide DNA nearest-neighbor thermodynamics".
# Proc. Natl. Acad. Sci. USA. 95 (4): 1460â5.
#
my %DP;
my %SN;
$SN{'A'} =1;
$SN{'G'} =2;
$SN{'T'} =3;
$SN{'C'} =4;
$DP{'AA'} = 0;
$DP{'TT'} = 0;
$DP{'AT'} = 1;
$DP{'TA'} = 2;
$DP{'CA'} = 3;
$DP{'AC'} = 3;
$DP{'GT'} = 4;
$DP{'TG'} = 4;
$DP{'CT'} = 5;
$DP{'TC'} = 5;
$DP{'GA'} = 6;
$DP{'AG'} = 5;
$DP{'CG'} = 7;
$DP{'GC'} = 8;
$DP{'GG'} = 9;
$DP{'CC'} = 9;
my $tDG=0;
my $tDH=0;
my $tDS=0;
my $fac=0;
my @DH=(-8.40, -6.50, -6.30, -7.40, -8.60, -6.10, -7.70, -10.10, -11.10, -6.70);
my @DS=(-23.60, -18.80, -18.50, -19.30, -23.00, -16.10, -20.30, -25.50, -28.40, -15.60);
my @DG=(-1.02, -0.73, -0.60, -1.38, -1.43, -1.16, -1.46, -2.09, -2.28, -1.77);
my @data;
my $seq;
my $ll;
my $naa;
my $Tm=0;
my $Tr=40.;
my $Ti=273.;
my $Tf=383.;
my $dt=2;
##########################################################################################
# Create the Tk GUI
##########################################################################################
my($pi, $x, $y) = (3.1415926, 0);
my $main = MainWindow->new(-background => 'yellow');
my $c = $main->Canvas(-width => 600, -height => 300, -bg => 'white' );
$c->pack(-side=>"top", -expand => 1, -fill => 'both');
my $text = $main->Scrolled('Text',
-relief => "sunken",
-borderwidth => 2,
-setgrid => "true",
-scrollbars => 'se' ,
-bg=>'cyan', -fg=>'black',
-height => 6
);
my $srly = $text -> Scrollbar(-background => 'yellow');
$text->pack(-side=>"top", -expand => 1, -fill => 'both');
$text->insert('end', "
DNA MELTING CURVE CALCULATOR \n
by Danilo Roccatano v1.0 \n ");
my $T1 = $main->Scale(-background => 'pink', -label=>"Temp. range :", -orient=>'h', -digit=>1, -from=>10, -to=>100, -variable=>\$Tr, -tickinterval=>5 );
$T1->pack(-side=>"top", -expand => 1, -fill => 'both');
my $T2l = $main->Label(-background => 'orange', -text => 'Enter Sequence:');
$T2l->pack(-side=>"top", -expand => 1, -fill => 'both');
my $S1 = $main->Entry(-textvariable=>\$seq, -validate => 'key', -validatecommand => sub {$_[1] =~ /[A|C|G|T]/i}, -invalidcommand => sub {$main->bell} );
$S1->pack(-side=>"top", -expand => 1, -fill => 'both');
my $b3= $main->Button(-text => 'RUN',-bg=>'green', -fg=>'black',
-command => sub{ runCalc (); } );
$b3->pack(-side=>"top", -expand => 1, -fill => 'both');
my $b4= $main->Button(-text => 'QUIT',-bg=>'green', -fg=>'black',
-command => sub{ $main->destroy(),exit; } );
$b4->pack(-side=>"top", -expand => 1, -fill => 'both');
MainLoop();
#####################################################################
#
# FUNCTIONS
#
#####################################################################
sub plot_gframe {
$c->delete('all');
#
# Create the upper graph
#
$Ti=$Tm-$Tr/2.0;
$Tf=$Tm+$Tr/2.0;
$dt=($Tf-$Ti)/490.;
# X -axis
$c->createLine(50, 250, 550, 250, -width => 2, -arrowshape => [ 10, 10, 10], -arrow => "last");
for ($x = 0; $x <= 490; $x += 25) {
$c->createLine(50+$x, 50, 50+$x, 260, -width => 1);
$c->createText(50+$x, 270, -fill => 'blue', -text => int($Ti+$x*$dt));
}
$c->createText(300, 285, -fill => 'blue', -text => 'T(K)');
# Y-axis
$c->createLine(50, 25, 50, 250, -width => 2, -arrowshape => [ 10, 10, 10], -arrow => "first" );
for (my $y = 0; $y <= 200; $y += 20) {
$c->createLine(40, 50+$y, 525, 50+$y, -width => 1);
$c->createText(30, 50+$y, -fill => 'blue', -text => 1.0-$y/200.);
}
$c->createText(50, 20, -fill => 'blue', -text => 'Diss.Fract.');
}
sub plot_mcurve {
my $s=200;
my $sy=1e10;
my $oy=250;
my $o=50;
my $sx=490./$Tr;
$c->createLine(50, 250, 50, 50, -width => 1);
my $xo=0;
my $yo=0;
my $dyo=0;
plot_gframe();
for (my $t = $Ti; $t <= $Tf; $t += $dt) {
# Calculate free energy
# dg = h - t * s + gplus
$x=$t-$Ti;
my $dG = $tDH - $t * $tDS;
my $A=$dG/$R;
# Calculate factor
$fac = exp(-$A / $t) * $Ce;
# Calculate fraction
$y = 1. - $fac / (1. + $fac);
$c->createLine($xo*$sx+$o, $oy-$yo*$s, $x*$sx+$o, $oy-$y*$s, -width => 1, -fill => 'blue');
$xo=$x;
$yo=$y;
}
}
sub runCalc {
$S1->cget('-text');
$seq = uc $seq;
$naa = length($seq);
my $RlC=$R*log($Ce);
my $cn = 0;
my $pp;
my $dpt;
$tDG = 0.0;
$tDH = 0.0;
$tDS = 0.0;
for (my $i = 0; $i < $naa-1; $i += 1) {
$cn++;
$dpt = substr($seq, $i, 2);
$pp=$DP{$dpt};
$tDG += $DG[$pp];
$tDH += $DH[$pp];
$tDS += $DS[$pp]/1000.;
}
#
# Add the correction values for the initial and final couples
#
$dpt = substr($seq, 0, 1);
$pp=$SN{$dpt};
if ($pp == 4 || $pp == 2) {
$tDG += 0.98;
};
if ($pp == 1 || $pp == 3) {
$tDG += 1.03 ;
};
#
# Check the chain symmetry
#
my $in = $seq;
$in =~ tr/ATCG/TAGC/;
my $rev=reverse$seq;
if ($in eq $rev) {
$tDG += 0.4;
}
#
# Calculte the melting temperature Tm
#
$Tm = $tDH/($tDS+$RlC);
#
# Print the results
#
$text->insert("end", "\nDNA SEQUENCE : $seq\n" );
$text->insert("end", "NUMBER OF NUCLEOTIDES : $naa\n" );
$text->insert("end", "\nDNA concentration (Ce/M): $Ce\n" );
$text->insert("end", "Dissociation Free energy (DG/Kcal/mol): $tDG\n" );
$text->insert("end", "Dissociation Enthalpy (DH/Kcal/mol): $tDH\n" );
$text->insert("end", "Dissociation Entropy (DS/Kcal/mol K): $tDS\n" );
$text->insert("end", "Melting Temeprature (Tm/Celsius): $Tm\n" );
$text->insert("end", "---------------------------------------------------\n" );
plot_mcurve();
}